Chan Lab:
Autophagy nutrient sensing



The ones who do the real work








Mahmud preparing some new mutant plasmids.
Ohood after taking care of her cell cultures.
Chinwe during an office chat.
Leon diligently washing up.
Laura, ready to talk about her poster.
Ben, from Hamburg for an ERASMUS exchange project.
Leon and I, before the run. 10k!
At the pub. Break from the lab
Bubble & squeak on special.
Ana, who travelled from Porto for an ERASMUS project with us.![]()
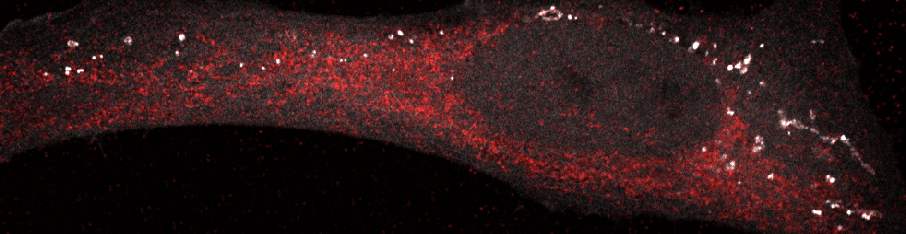
Our subject of study.
We work with human and mouse cells to learn about biology and medicine.
We have a range of normal and cancer cells frozen away. Most of our cell models are immortalized, which provides us with consistent and renewable biological systems for study.
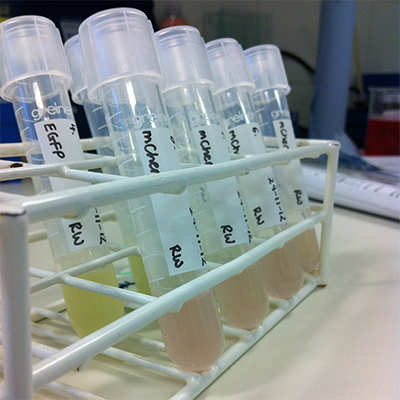
To study cell biology, we rely heavily on tagged proteins that we can visualize.
Here, we prepare new GFP (green fluorescent protein) and RFP-(red) tagged plasmids using bacteria as an intermediate host. The GFP and RFP are so strong they colour the bacteria.
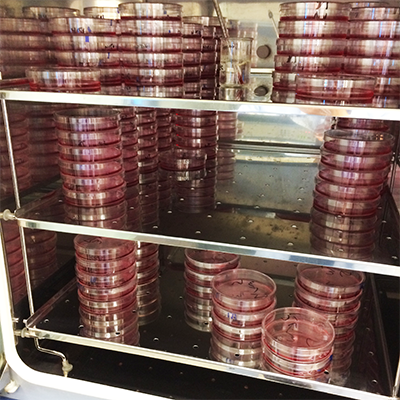
We aim to find new regulators of our favourite pathway, autophagy.
Here, we use a genetic screen to find new mutants. In this collection of dishes, there are mutations in each of the 20,000 genes in the human genome.
We then select for the mutant phenotype that we want.
Once we find new genes, then the real grind work starts with smaller experiments testing our hypotheses.
We aim to find out how genes regulate signalling and cell fate.
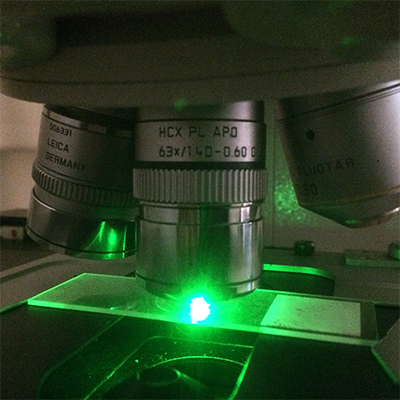
One of our preferred approaches once we focus down is to visualize spatial processes going on inside cells.
For this, we use laser scanning confocal microscopy.
We learn a lot from studying membrane structures inside cells.
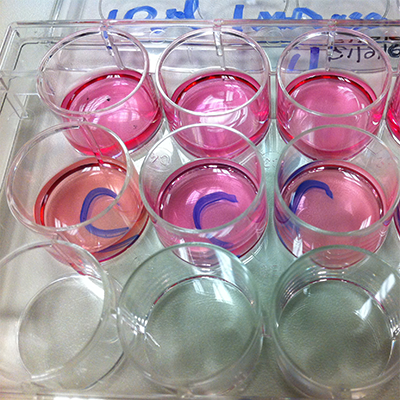
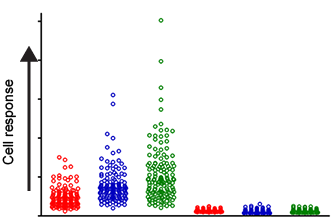
We measure responses across cell populations which enables us to differentiate between strong and subtle effects.
If we find a result that supports our hypothesis, success!
Stop at the pub to celebrate, before starting again the next day.
L M H L M H
Normal Mutant
I am just a child who has never grown up. I still keep asking these 'how' and 'why' questions. Occasionally, I find an answer.
Stephen Hawking